Understanding the mechanisms of plasmid replication can be useful to develop strategies against bacterial antibiotics resistance
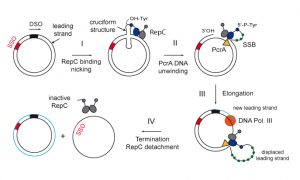
RepC is a 38-kDa homodimeric protein encoded by the pT181 plasmid of Staphylococcus aureus. It binds to a specific DNA sequence contained within the double-stranded origin of replication (dso) of the plasmid and creates a nick (step I). The nicking site is contained within an inverted repeat that is likely to form a cruciform structure. RepC nicks the leading strand and becomes covalently bound to its 5´-end via a phosphotyrosine linkage. Then, PcrA helicase loads onto the short length of accessible single-stranded DNA (ssDNA) (step II). The free 3´-end at the nick is used for extension synthesis where PcrA, DNA polymerase III, and single-strand DNA binding protein (SSB) coordinate their activities at the origin to replicate the entire plasmid (step III). Replication of DNA by DNA polymerase III proceeds until the plasmid is fully unwound and a new leading strand has been fully synthesized. During elongation, the displaced leading strand forms a single-stranded loop that is protected by SSB proteins from nuclease degradation. SSB protein also greatly stimulates the incorporation of nucleotides in vitro by blocking non-productive binding of the DNA polymerase to ssDNA. At the termination step, the other monomer of RepC protein cleaves the displaced ssDNA at the regenerated nick site at the junction of the old and newly synthesized leading strands. Next, in a process that is not fully understood, the nick is sealed by RepC leaving a relaxed, closed circular double-stranded DNA (dsDNA) containing the newly synthesized leading strand (step IV). The relaxed DNA is supercoiled (SC) by DNA gyrase while the RepC protein is released from the DNA in an inactive form, covalently bound to a short DNA fragment. The displaced single leading strand is then converted to the double-stranded form by using the single strand origin of replication (sso), and requires RNA polymerase for primer synthesis and DNA polymerases I and III for replication.
Want to learn more?
Pastrana*, Carrasco* et al. Nucleic Acids Research 44(18), 8885-8896 (2016). Force and twist dependence of RepC nicking activity on torsionally-constrained DNA molecules. LINK
