DNA crookedness regulates DNA mechanical properties at short length scales
A. Marin-Gonzalez*, J.G. Vilhena*,F. Moreno-Herrero† and R. Perez†
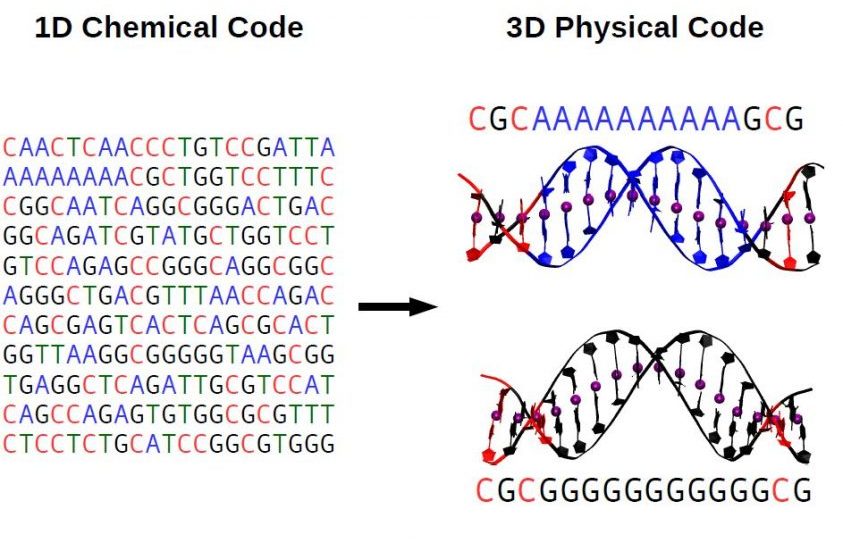
Sequence-dependent DNA conformation and flexibility play a fundamental role in the specificity of DNA-protein interactions. Here we quantify the DNA crookedness: a sequence-dependent deformation of DNA that consists of periodic bends of the base pair centers chain. Using extensive 100μs-long, all-atom molecular dynamics simulations, we found that DNA crookedness and its associated flexibility are bijective, which unveils a one-to-one relation between DNA structure and dynamics. This allowed us to build a predictive model to compute the stretch moduli of different DNA sequences from solely their structure. Sequences with very little crookedness show extremely high stretching stiffness and have been previously shown to form unstable nucleosomes and promote gene expression. Interestingly, the crookedness can be tailored by epigenetic modifications, known to affect gene expression. Our results rationalize the idea that the DNA sequence is not only a chemical code, but also a physical one that allows finely regulating its mechanical properties and, possibly, its 3D arrangement inside the cell.